 Intellectual labels are always a tricky business, necessary for talking about ideas and suggesting that a theorist is in a particular ideological neighborhood. Yet, they can drag along so much baggage that they become self-defeating, evoking instant resistance or inevitable misinterpretation if poorly used. In the best of cases, they can help to create a clear identity for innovative work in an academic field, speeding the effort to carve out a space for ideas in a cluttered terrain of thought. Deployed well, they can help to clarify and orient us; applied clumsily, they become intellectual invective, prematurely close off discussion or debate, and substitute labeling for thinking.
Intellectual labels are always a tricky business, necessary for talking about ideas and suggesting that a theorist is in a particular ideological neighborhood. Yet, they can drag along so much baggage that they become self-defeating, evoking instant resistance or inevitable misinterpretation if poorly used. In the best of cases, they can help to create a clear identity for innovative work in an academic field, speeding the effort to carve out a space for ideas in a cluttered terrain of thought. Deployed well, they can help to clarify and orient us; applied clumsily, they become intellectual invective, prematurely close off discussion or debate, and substitute labeling for thinking.
Today, I want to write briefly about ‘neuroanthropology’ as a badge, but spend more time on ‘neuroconstructivism,’ as it’s a term that sometimes gets associated with the sort of research and thinking that we are advocating here at Neuroanthropology.net. In a sense, this piece is written for non-anthropologists, to help them understand why they might get a really strange reaction from an anthropologist colleague if they start talking excitedly about new ‘neuroconstructivist’ perspectives.
We’ve obviously decided that ‘neuroanthropology’ is one of the labels that we find helpful. We stand by the neologism, even though some of our readers have described our choice of terms ‘deplorable,’ and we’ve sometimes had to struggle against the term’s use elsewhere. For example, Oliver Sachs, the wonderful chronicler of the lived worlds of people with severe brain lesions, often calls himself a ‘neuroanthropologist,’ as Jovan Maud at Culture Matters pointed out to me and Daniel highlights in a recent, more thorough post on the relation of what we’re doing to what Sachs has done (see also Neuroanthropology).
For Sachs, the emphasis is on the ‘neuro-’ with the ‘anthropologist’ part of the term referring more to the genre or style in which he writes and his humanistic concern for the sufferer’s-eye-view in neuropathology. That is, Sachs uses the term to inflect ‘neuro-’ with a sympathy for the sufferer’s own perceptions, what in anthropology we call an ‘emic’ or ‘experience-near’ perspective. I’m not sure when he started using the term, but one of his subjects, the brilliant and autistic Temple Graddin, used it to describe her sense of being an outsider in everyday interactions, like an ‘anthropologist on Mars.’
Unlike anthropologists, Sachs doesn’t really consider the range of normal (non-‘pathological’, with all the challenges that this label brings along). He also tends to study cases in isolation, without much of a focus on the social relations, cultural influences, and interplay between factors like environment, education, inequality, ideology, and the like that is a hallmark of contemporary anthropology. So although I’m very happy to be associated in any way with the remarkable Oliver Sachs, I would say that we are putting the ‘neuro-’ in the ‘anthropology’ to get neuroanthropology, perhaps working in the opposite direction and on slightly different terrain than the good Dr. Sachs. Still, if there’s any guilt by association, he’s good company to keep.
Still, the fact that we’re trying to bring the ‘neuro-‘ to ‘anthropology’ helps me to explain why I don’t like the term ‘neuroconstructivist’ even though I like nearly everything about neuroconstructivists themselves.
Okay, so I admit I was over-reacting…
What really got me thinking about intellectual labels and this issue was a one-line remark that pissed me off a while back when I first read it. Oliver Morin, who writes great stuff for the Cognitive and Culture Institute weblog, posted a veeeeeery short comment on a piece that I wrote on an article by Andy Clark and William Wheeler. Morin wrote: ‘Neuroanthropology dwells at some length on a paper published by Andy Clark and William Wheeler in the latest PTRS special issue. It is basically a celebration of neuroconstructivism. [emphasis added]‘
For a while, that brief comment by Morin kept irritating me, but I knew his work enough to know that he is a sophisticated thinker, not one to simply consign what I found to be a really interesting article by Clark and Wheeler with a shallow label (I know, I know – it was only a one-line description. A propensity to mull things over extravagantly is likely an adaptive trait for an academic.). I had to dig a bit into the literature to find what specifically was meant by ‘neuroconstructivism,’ in the process running into a bibliography that included a number of my favourite thinkers on subjects like emergence, cognitive development, developmental psychology, and evolutionary theory. It also occurred to me that there was likely to be a serious translation problem when the term ‘neuroconstructivist’ crossed over into anthropology if it set my teeth on edge, and I’m both reasonably neuro-savvy and trust the person who used the term.
Neuroconstructivism: not just another constructionism
The term ‘neuroconstructivism’ bothered me at first because of its close proximity to ‘social constructionism’ and ‘cultural constructionism,’ and, by extension, ‘deconstruction.’ (Although some proponents draw a distinction between social constructionism and social constructivism, I’m going to ignore that distinction because it’s inconvenient for my argument, at best. Besides, it’s already an overly-long blog post.)
Various forms of social constructionism have encouraged contemporary cultural anthropologists to focus on the degree to which important collective social products such as ideologies, narratives, languages, imaginaries, and discourse are social fictions. The short definition of ‘social constructionism’ (and it is anything but ‘short’ on internal disagreements and variations) is that social constructs, the products of human interaction – whether they be ideas, material artifacts, or practices – persuade socialized actors of their reality and independence from the actors’ perception of these fictions. Social constructs appear to be objective reality.
Social constructionists focus on the arbitrariness and implications of constructs, even if actors take them to be accurate representations of reality often precisely because the users of constructs don’t recognize that they are not operating with simple reflections of a reality external to culture. For social constructionists, these social fictions can then be seen to have all sorts of consequences, some of them quite material and even detrimental to those who believe in the fictions.
Clearly, certain facts or objects, however persuasive, are culturally specific and sufficiently autonomous from brute reality that it’s hard to disagree that they are social constructs. That is, it’s very hard to argue that, without customs, belief, and social consensus, entities like money, law, language, nations, reputations, and the like have much existence prior to human conviction that they exist. As we’ve seen in the recent financial crisis, some social constructs are quite precarious; rattle the markets a little, and ‘assets’ like mortgaged-backed securities or financial derivatives, even some ‘corporations,’ turn out to be shaky fictions indeed (‘Wasn’t there just a Fortune 500 corporation here a minute ago?’).
Observers sometimes draw a distinction between ‘strong’ and ‘week’ social and cultural constructionism. Strong varieties of social constructivism, in the work of philosophers like Jean-François Lyotard and some anthropologists, argue that reality itself is a product of social consensus, that we interact, not with a reality external to us, but with social fictions that we have created. In truth, strong social constructionists are rare, although a fair bit of contemporary cultural anthropology has a tendency to look like strong social constructionism because writers often skirt the thorny problem of how ‘real’ native concepts are and don’t really grapple with some of the materiality of cultural life.
For example, many anthropologists are much more comfortable talking about indigenous concepts of illness and healing than asking hard questions about causes, consequences of different therapies, and mechanisms for understanding how they work. Since we started Neuroanthropology.net, I’ve actually grown more sympathetic to this approach because I now better recognize how deep the theoretical water is and how difficult the empirical questions are if you want to think about the relationship of cultural concepts to things like biological causation.
But most social and cultural anthropologists, although they may write extensively about social constructions, are not strong social constructionists. They acknowledge that material reality is obdurate, that biology plays a part, that genes influence development, that language is both social creation and response to an objective world, but they tend not to focus on these parts of the mangle of everyday experience because the field has spent so much of its time trying to disabuse other people, academics and civilians alike, that reality wasn’t as simple and objective as they might think. That is, I suspect that, although they spend much of their professional lives working in constructionism, anthropologists don’t wander around in their off hours spiraling in existential contemplation of the fictions that surround them.
Looks like a neuroconstructionist, quacks like a neuroconstructionist…
To neuroscientists, ‘neuroconstructionism’ is an analytical perspective that focuses on the emergence of neurological abilities out of less complex neurological abilities. As Westermann and colleagues (2007:75) describe, many developmental psychologists tends to explore ‘children’s abilities at specific ages without devoting equal attention to the question of the mechanisms by which these abilities unfold and change over time.’ In constrast, neuroconstructivism tries to ‘link the observed abilities of infants and children at different ages into one developmental trajectory’ (ibid.).
Neuroconstructivism is also discussed in the two-volume collection of that name (Mareschal, Johnson, Sirois, Spratling, Thomas & Westermann 2007; Mareschal, Sirois, Westermann, and Johnson, eds. 2007).
One of the crucial characteristics of neuroconstructivism is the recognition that neural activity, affected by the environment, behaviour patterns, and a host of other influences, effects change in neural architecture: ‘cognitive processing itself shapes the neural networks that are responsible for this processing in the first place’ (Westermann et al. 2007:75). Acknowledging the fact that no specific level of analysis is determining the others (neither gene determining behaviour, nor behavioural independence from biology), means treating cognitive and neural levels of description as inseparable; neither can be studied in isolation from the other. As Westermann and colleagues (ibid.:76) write, neuroconstructivism calls, not for reduction of phenomena to neurological correlates, but for ‘consistency between the neural and cognitive levels in characterizing developmental trajectories.’
Westermann and his colleagues (ibid.) highlight six levels of interrelated phenomena, but I don’t think they have any intention of suggesting these are exhaustive:
1) insights from epigenesis on the probabilistic relationship between gene and expression, including the impact of behaviour on gene expression;
2) how experience affects the development of different neural structures;
3) how regional brain specialization emerges through interaction among brain regions;
4) how bodily experience affects cognitive development;
5) how the child’s own activity and active pursuit of its own goals affect its cognitive development; and
6) how social environments affect the developing child.
There’s so many problems with the old organism-environment dichotomy that it’s a bit of dead horse abuse to flog it anymore, but one thing that comes out of Westermann et al.’s discussion is that, depending on the scale of the analysis, what is the ‘environment’ can be radically different. For the gene, the cell is the environment; for the individual organ, such as a brain region, the body is the environment; but at the scale of the organism as a whole, these smaller scale ‘environments’ are, of course, the organism which is nested in still greater concentric environments. Westermann and colleagues (ibid.:77) write, ‘As the brain is embedded in a body (embodiment [note: not what anthropologists mean by ‘embodiment’]), so an individual functional brain region is embedded in a brain where it co-develops with other brain regions.’
Westermann and colleagues offer the following figure to outline some of the relations in embodiment:
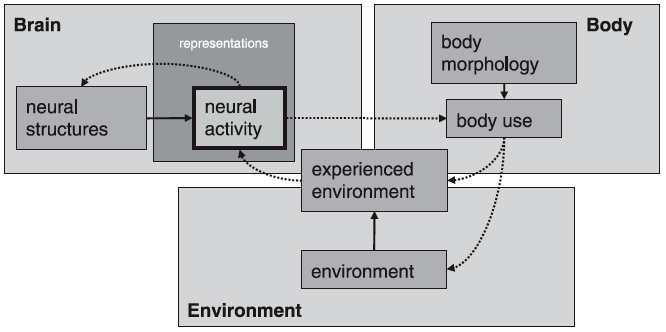
(By the way, I’ll be back to this flowchart as I’m working on a bit of a piece on Westermann et al.’s nifty graphics.)
In a sense, neuroconstructivism is a synthesis of some of the most promising integrative research that we have been exploring on Neuroanthropology.net. Although I could cite a number of passages to highlight the type of thinking that the neuroconstructivist perspective offers, I’ll just leave you with one from Westermann et al.’s discussion of ‘interactions between constraints’:
Interactions with a social environment have effects on both neural development and on the expression of genes (Eisenberg, 1995). These effects can either be mediated through direct experience with the environment or through altered caregiver behaviour in a specific environment (Sale, Putignano, Candedda, Landi, Cirulli, Berardi & Maffei, 2004).
That is, social environments affect biological development through several different mechanisms, some of them direct environmental contact and some of them indirect through the environment’s effect on the caregiver.
What’s in the name, neuroconstructionist
This is why I have reservations with Olivier Morin’s use of the label ‘neuroconstructivism,’ even though he’s spot on and has every right to label accurately (in other words, he’s right, and I still have a problem). After the AAA conference and some of the conversations I’ve had, I’m not content with the term, not because of what it means to neuroscientists but because of how it will likely be understood by anthropologists. The gap between our field’s understanding of ‘constructionism’ and this conception of ‘neuroconstructivism’ is huge and yet it may be invisible to anthropologists.
I can almost hear my colleagues right now: ‘Neuroconstructivism’? What would that be? ‘Constructivism’ plus ‘neuro-‘, I suppose. Sounds like ‘social constructionism’….
Then, there’s a moment of hesitation as the anthropological reader tries to decide whether to attack (‘They’re saying our neurons construct us!’) or to prematurely agree with an incorrect understanding of what we’re about (‘Oh yeah, neurons are a social construct; that’ll really piss off the neurologists. I’m down with that!’). Especially when what follows is likely to be complex material on multiple scales, some of which might be unfamiliar to an anthropologist, I’m afraid that the response is liable to be either entrenched, knee-jerk resistance to ‘biological reduction’ (it’s obviously not reductionist from even the short description) or overly-quick acceptance without digesting the real significance of what’s being argued (this is not your retired thesis supervisor’s ‘constructionism’).
Either way, we don’t get what we want, which is really a more subtle discussion of the interaction between different scale processes, from the cellular and neural to the developmental, social and cultural. That is, I fear that the closeness of the label ‘neuroconstructivist’ to a set of older terms might signal battle… errr, business as usual.
In anthropology the label ‘x-constructivist’ (following the examples of ‘cultural constructionist’ and ‘social constructivist’) indicates that the person being tarred with the brush places the overwhelming emphasis in any model on the ‘x-‘ against ‘y-innatist’ or ‘z-determinist.’ That is, there are two poles in some anthropological thinking, one ‘constructivist’ that emphasizes ongoing influences (culture, social interaction, upbringing, environment…) and the other ‘determinist’ that emphasizes innate traits (genetic, biological…). Part of my own strategy as a neuroanthropologist is to try to avoid saying things that let my readers off the hook, allowing them to engage in old-fashioned, one-sided, reductionist thinking that has made biocultural synthesis hard to imagine.
I don’t think Morin has ANY intention of dragging this tired constructivist v. determinist dichotomy into the new territory, but this may be a difference between European and American anthropology. Or maybe Morin is sufficiently recovered from the Constructionist-Determinist War and I’m still suffering from the lingering fear that the conflict will re-ignite. (‘The horror!’) The terms we use have to be carefully chosen, and translation between fields is sometimes hairy, especially if we see a term and don’t realize it even needs translation.
For example, one term that I’ve brought up before is ‘representation’; in the neurosciences, the term means the physiological neural correlates that produce a percept, concept, or other neural activity. Westerman and colleagues (2007:75), for example, offer the following: ‘Representations are here defined as neural activation patterns in the brain that contribute to adaptive behaviour in the environment.’ The term is debated, in some quarters of the brain sciences, though it is widely accepted, because it may imply a degree of stability, fixity or clear structural location that some theorists argue is unwarranted.
My problem with ‘representation’ like my problem with ‘neuroconstructivist’ is that I’m worried about how they will cross over into an anthropological audience; in particular, I’m concerned that these terms will blunt the potentially positive theoretical challenges that research in the brain sciences might pose for anthropological theory. Anthropologists talk about ‘representations’ and ‘construction’ a lot, but they do not mean the neural activation pattern that contributes to adaptive behaviour nor do they mean a sensitivity to the complex unfolding of organism-environment interactions in developmental timeframes (see, for example, the discussion in Beyond Bourdieu’s ‘body’ — giving too much credit?).
I personally prefer ‘neurocultivationist’ to ‘neuroconstructionist,’ but I feel like we’ve likely used up our neologism allotment for the year here at Neuroanthropology, so I probably won’t get my way. ‘Cultivation’ has a bit more organic implications, a bit less of a metaphoric extension to teleological or plan-driven development. One ‘constructs’ a house from a plan, digging out a foundation with a backhoe and tearing out trees to realize a pre-existing vision; one must work with nature and over time to ‘cultivate’ a garden, never sure how or even if everything will grow.
So even though Morin is right, and I was wrong to get testy about being called a ‘neuroconstructivist,’ I still won’t use the term around anthropologists too much. I don’t want them to think they actually know what I mean.
Credits: Graphic from iStockphoto.
References
Eisenberg, Leon. 1995. The social construction of the human brain. American Journal of Psychiatry 152 (11): 1563–1575. (abstract, pdf available here)
Mareschal, Denis, Mark Johnson, Sylvain Sirois, and Michael Spratling, Michael Thomas, and Gert Westermann. 2007. Neuroconstructivism: How the brain constructs cognition. Oxford: Oxford University Press. (Amazon)
Mareschal, Denis, Sylvain Sirois, Ger Westermann, and Mark Johnson, eds. 2007. Neuroconstructivism vol II: Perspectives and prospects. Oxford: Oxford University Press.
Sale, Alessandro, Elena Putignano, Laura Cancedda, Silvia Landi, Francesca Cirulli, Nicoletta Berardi, and Lamberto Maffei. 2004. Enriched environment and acceleration of visual system development. Neuropharmacology 47(5): 649–660. (abstract)
Westermann, Gert, Denis Mareschal, Mark H. Johnson, Sylvain Sirois, Michael W. Spratling and Michael S.C. Thomas. 2007. Neuroconstructivism. Developmental Science 10(1): 75–83. doi: 10.1111/j.1467-7687.2007.00567.x (pdf available here)

 Stumble It!
Stumble It!
As usual, a great, stimulating post.
A great post, but I’m left wondering exactly what neuroconstructivism is. At least as you’ve outlined it, it seems like it would be very hard to not be a neuroconstructivist, if you were serious about doing neuroscience. The “six points” of Westermann et al sound almost like a set of commandments : Thou Shalt Not Forget That Social Environments Affect the Developing Child. Which is perfectly true, but if that kind of thing is all neuroconstructivism is, then it doesn’t seem like a theory per se. More of a movement with the goal of making sure that neuroscientists don’t slip into lazy cod-reductionism.
Good point, Mr. Neuroskeptic: I don’t really think neuroconstructivism is a ‘theory per se,’ but that’s probably something I should state more clearly. You’re probably right — or at least I hope you are — that a neuroscientist would have to be a neuroconstructivist of some sort if he or she were studying any sort of neural processes over time (the only way to study something like learning, trauma recovery, or plasticity, for example).
But I do think that this ‘perspective’ (rather than theory, let’s say) is not always the starting point of a lot of other theorizing grounded, or allegedly grounded, in neurological evidence. That is, I think that a lot of neural-based theorizing takes a static brain or a ‘finished’ brain as a starting point, rather than really integrating the constructivist dimensions, the emergent nature of neural structures, into the heart and soul of the theorizing. For example, I think a lot of modularity thinking is incompatible with a basic recognition of the issues that neuroconstructivists are pointing out. Likewise, some approaches to evolutionary psychology are also harder to defend if emergence in brain structure is taken seriously. It’s not just ‘environment affects child’ but the many levels on which these structures are emerging — intercellular relations, modularization (specialization of tissue over time), necessity of sensory input for neural development, as well as the more traditional ‘environmental’ concerns such as parent-child relations, interpersonal environment, language socialization and the like.
That is, I think neuroconstructivism points to a ‘environment-all-the-way-down’ set of concerns, treating ‘environment’ on a spectrum of scales, that forces a consideration of developmental processes. I also think that the Westermann et al. article is probably not as interesting as the books (which I have ordered for our library) because at the general level they’re writing, these points seem very abstract, much like West-Eberhardt’s book on developmental plasticity is especially formidable because of the staggering variety and depth of the examples, rather than the complexity of the central argument.
Readers may be interested in Dr. Susan Sheridan’s “Scribble Hypothesis” in which she posits that the very young child, while scribbling, is actually building brain cells. This is what neurocontructivism means to me, as an art educator.
drawingwriting.com is her website.